- Home
- Details
- Registry
- Blog
- RSVP
- Como descargar el gta san andreas para pc
- Mason hamlin model 50
- Visual studio 2012 express free download
- Slobodan trkulja banski dvori
- Apache air assault code
- Android infinity box
- Contax g2 with 28mm
- Mestrenova nmr peaks prediction
- Linux support for anker usb hub with ethernet
- J tag for xbox 360
- Ninja village best layout
- Gaming mouse razer deathadder elite
a certain institution they have been measured at. The spectrum categories, on the other hand, should refer to the spectrum, e.
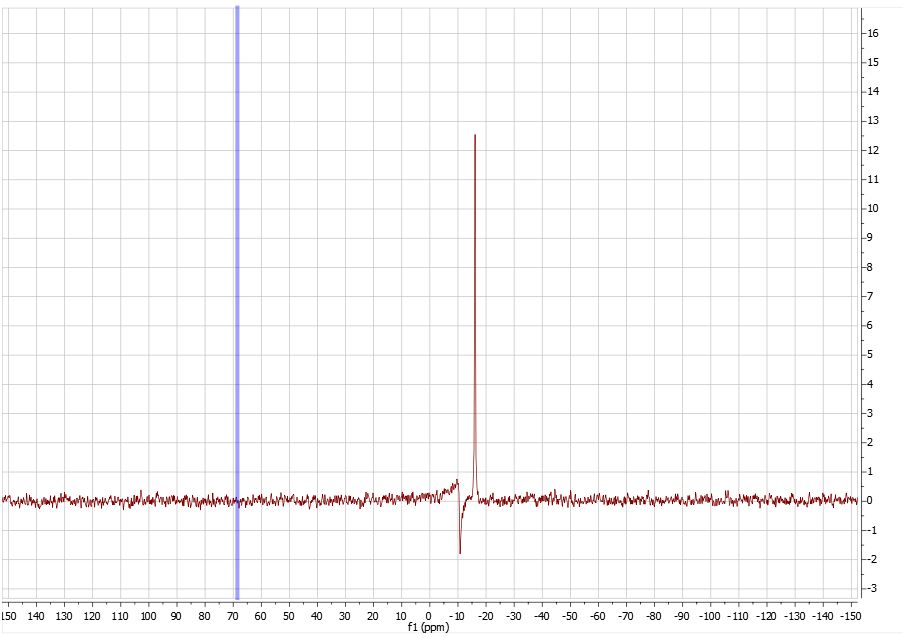
Molecule keywords should be compund-specific, e.
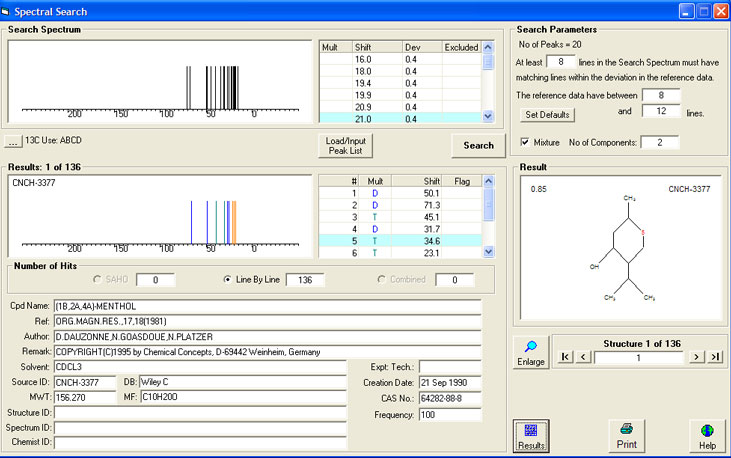
Further more, you can give keywords for the molecule and categories for the spectrum - either existing ones or new ones. You can also enter literature and web pages about the spectrum. From time to time IUPAC and index names for molecules in nmrshiftdb2 will be auto-created, so you do not need to enter them. Then enter the chemical names, any links to web-literature about the molecule and the CASNumber and autonom name of the molecule, if known (you do not need to do that if the molecule is already in the database). If you have not entered the shifts before this step, you can them enter here directly.Īdd miscallenous data: Please choose first if the spectrum is measured or calculated. You can also enter multiplicities for atoms (except for proton spectra, the default is calculated by the proton count of an atom) and coupling constants for every atom-atom combination. If you have entered only one shift, it gets assigned to all atoms automatically and you may skip this step. If you forgot to submit a signal, finish this step and go to "Add shifts" and then "Do assignments" again. After you assigned all signals, press the "Submit assignments" button. Shifts not assigned can be deleted in a later step or kept as unasssigned - this is also a way to get rid of mistyped shifts. You can also edit an existing set by changing some identifiers. "Standard" means the numbers as they appear in the structure drawing. If you enter a spectrum for an existing structure you can chose an existing set of identifiers below the structure diagram. You can also use this to renumber the atoms by using numbers as identifiers. If there is no identifier for an atom the number from the "Atom number" column will be used. You should make sure identifiers are unique. In the "Atom identifier" column you can enter identifiers for the atoms (any character allowed) which will be used instead of the standard numbers. Blue figures indicate that your value is inside this range, red means outside. In some rare cases the predicted value will be identical with one of the limits of the range. Additionally there is a range for the signal given. These predictions are calculated based on HOSE codes, as in the prediction function. These peaks will not be processed in any way, but saved as given.ĭo assignment (for standard 1D spectra only): Here you have got one line per atom, showing a drop-down box for the shifts and the expected range for the shift. The third gives the coupling constant and is optional. The first two figures are the values on the two axis. If you submit, this peak list will replace the automatic peak list.įor 2D spectra the format is slight different: Then the average of all peaks in this range will be calculated and added to the peak list on the right. You need to left click on the field and go over some peaks and release the mouse. Most likely, these will contain multiplets, noise etc. You will be presented with an applet containing all individual peaks.
#Mestrenova nmr peaks prediction manual#
If the spectrum type is 1H, you need to perform a partly manual peak picking. If the spectrum type (nucleus) is in the file, it will be used, else you also need to choose the spectrum type. If the file contains a peak table, this will be used, if not, a peak picking is performed. You can also upload a jcamp file by hitting "Browse." and choosing the file. You can add additional signals later by the same steps. After you input the spectrum, click the "Submit signals". They will be scaled to be in a range from 0 to 1, if they are not. For 1D spectra you need to give one signal per line, shift and intensity separated by. Set spectrum type: Here you enter the type of the spectrum (13C, 1H etc.) you want to submit.Īdd shifts: Here you enter the signals of your spectrum. If you want to submit more than one spectrum you can repeat the upload after submitting a spectrum. If there is more than one spectrum in the files you can choose which one you want to submit in the next step ("Set spectrum type"). From the sd file the shift lists will also be read. The structure and custom numbering, if in the file, will be read. You can upload mol and sd files from MNova (sd files only work in MNova 11 or higher) or mol files from ChemDraw with labels via the "Upload" facility below the structure editor. If you want to change the structure later, just hit "Enter molecule" again. Your structures are collected in the structures history. After you drew or imported your structure, submit it by clicking the blue "Submit molecule" button. You do not need to (but you may) draw implicit hydrogens.
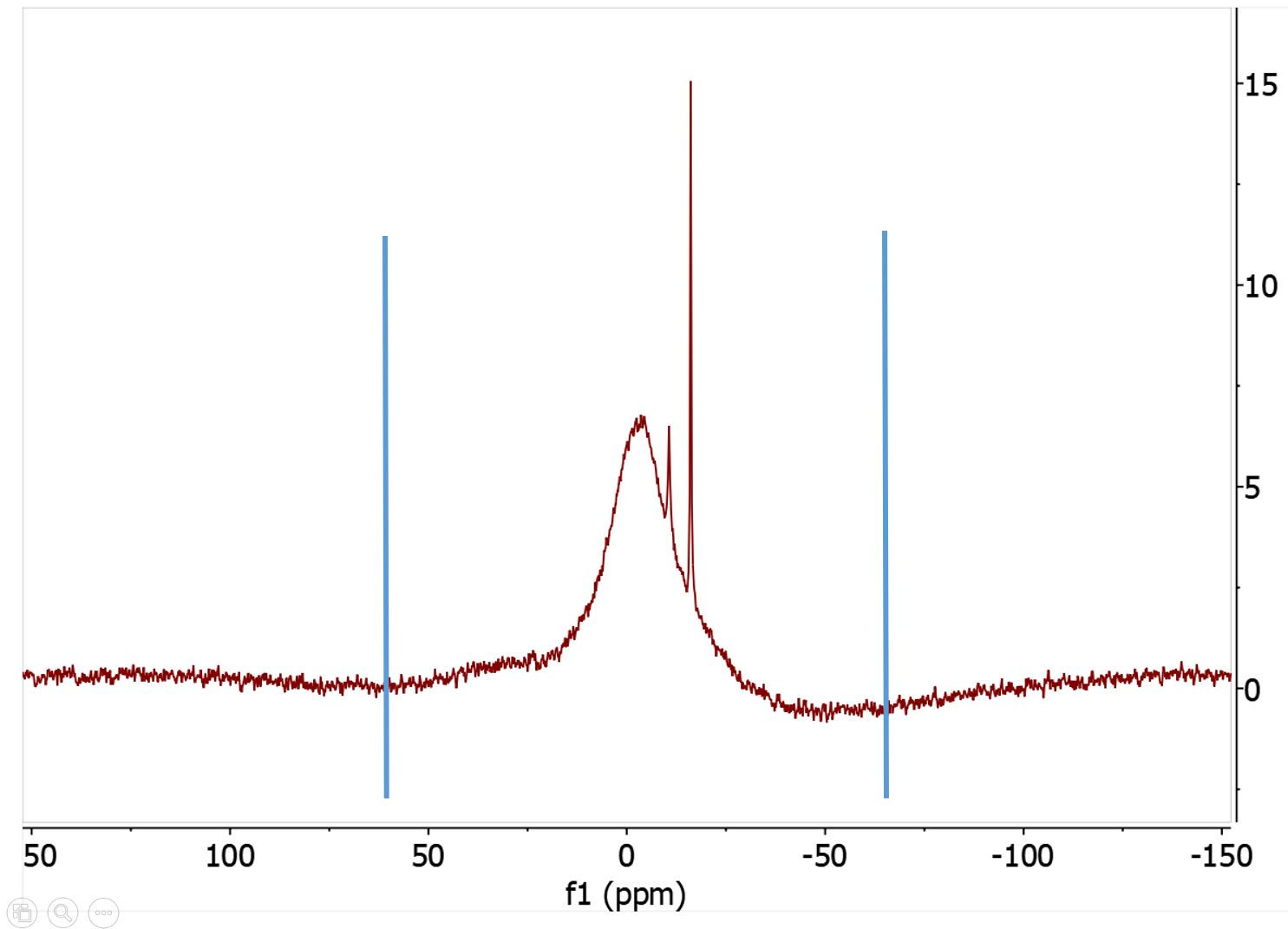
You can also import a file by Copy & Paste via the "Import text by copy &paste button". You can then use the JChemPaint applet to interactivly paint structues.
- Home
- Details
- Registry
- Blog
- RSVP
- Como descargar el gta san andreas para pc
- Mason hamlin model 50
- Visual studio 2012 express free download
- Slobodan trkulja banski dvori
- Apache air assault code
- Android infinity box
- Contax g2 with 28mm
- Mestrenova nmr peaks prediction
- Linux support for anker usb hub with ethernet
- J tag for xbox 360
- Ninja village best layout
- Gaming mouse razer deathadder elite
